Church Evolution Laboratory

Department of Biology, New York University
![]()
Research
Our research group studies the evolutionary process of diversification. We combine field work, lab work, and computational analysis to understand how variation emerges across biological scales, from genes to cells, organisms, and populations.
This research spans invertebrate systems, including projects on Hawaiian fruit flies as a model for evolutionary development, sailing jellyfish found at the ocean surface, and life-history trade-offs in insect eggs and ovaries.
Because our work often involves new technologies and tools for collecting data across species, we also develop and release open-source software for analyzing and comparing large datasets.
Hawaiian Drosophila: model clade for evo-devo
Millions of years ago, Drosophila fruit flies found their way to the Hawaiian islands. Now there are many hundreds of species found on Hawaii and nowhere else in the world.
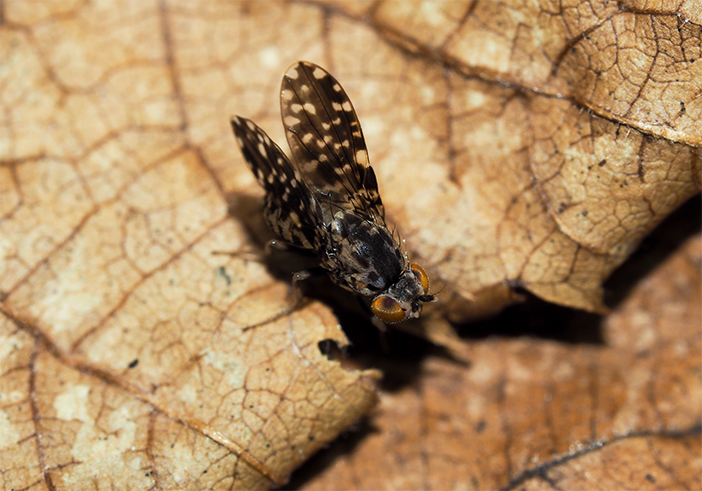 Adult Hawaiian fly Drosophila picticornis
Adult Hawaiian fly Drosophila picticornis
Given their close relationship to model species like Drosophila melanogaster, this group is primed to be a model clade for evo-devo. In our research, we combine next-gen sequencing and phylogenetic comparative methods to uncover their evolutionary history.
The central question of our research is: What drives diversification? Hawaiian Drosophila present an ideal system to study this question, given their spectacular diversity in morphology, behavior, and ecology, generated over a relatively short evolutionary timescale.
See the Hawaiian Drosophila Genomes Project for our collaborative initiative to sequence and compare the genomes of all ~1,000 endemic Hawaiian fly species.
Evolutionary inference for developmental data
We develop new tools for evolutionary analysis of gene expression data from single-cell sequencing and other RNA-seq technologies. This work combines mathematics, bioinformatics, and developmental and evolutionary biology to build a new analytical framework.
We advocate for analyzing data in the context of robust null hypotheses that describe how much variation we expect to observe when we compare across species.
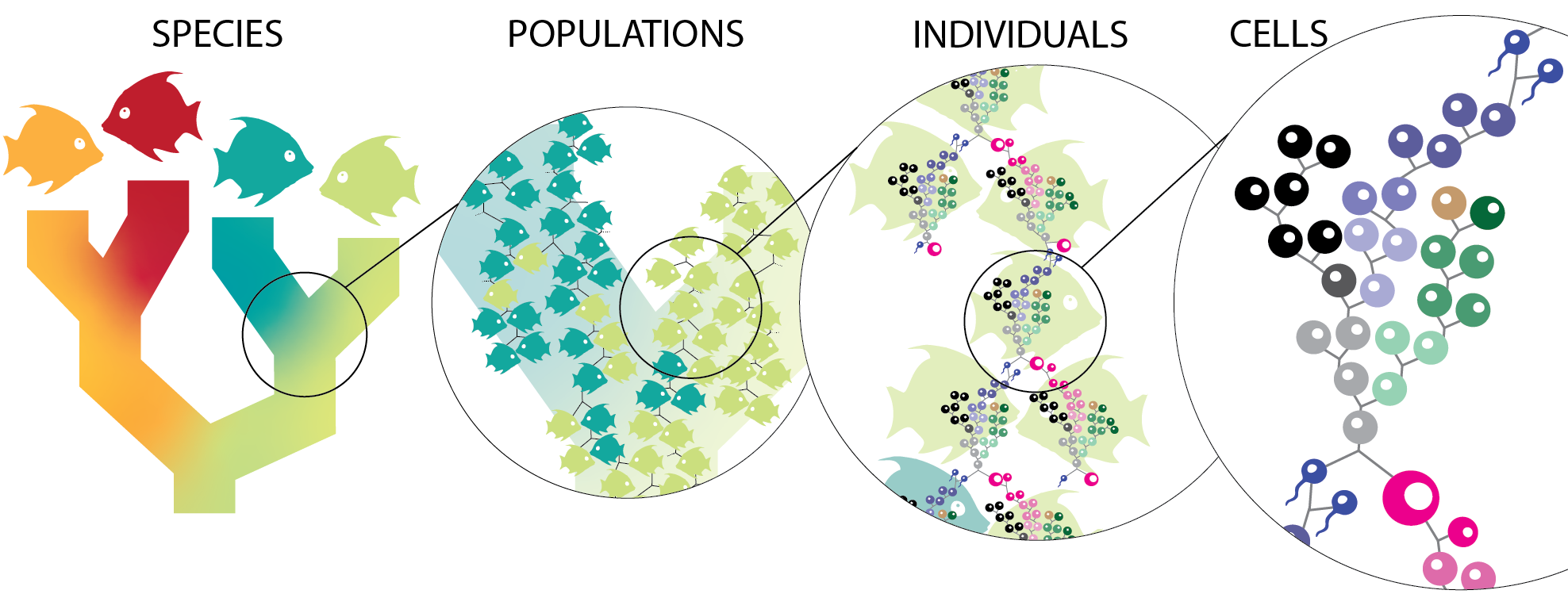
The objective of this work is to infer and interpret the cellular tree of life. This tree unites species phylogenies, with population maps, geneologies of individuals, and down to cell lineages within a single individual.
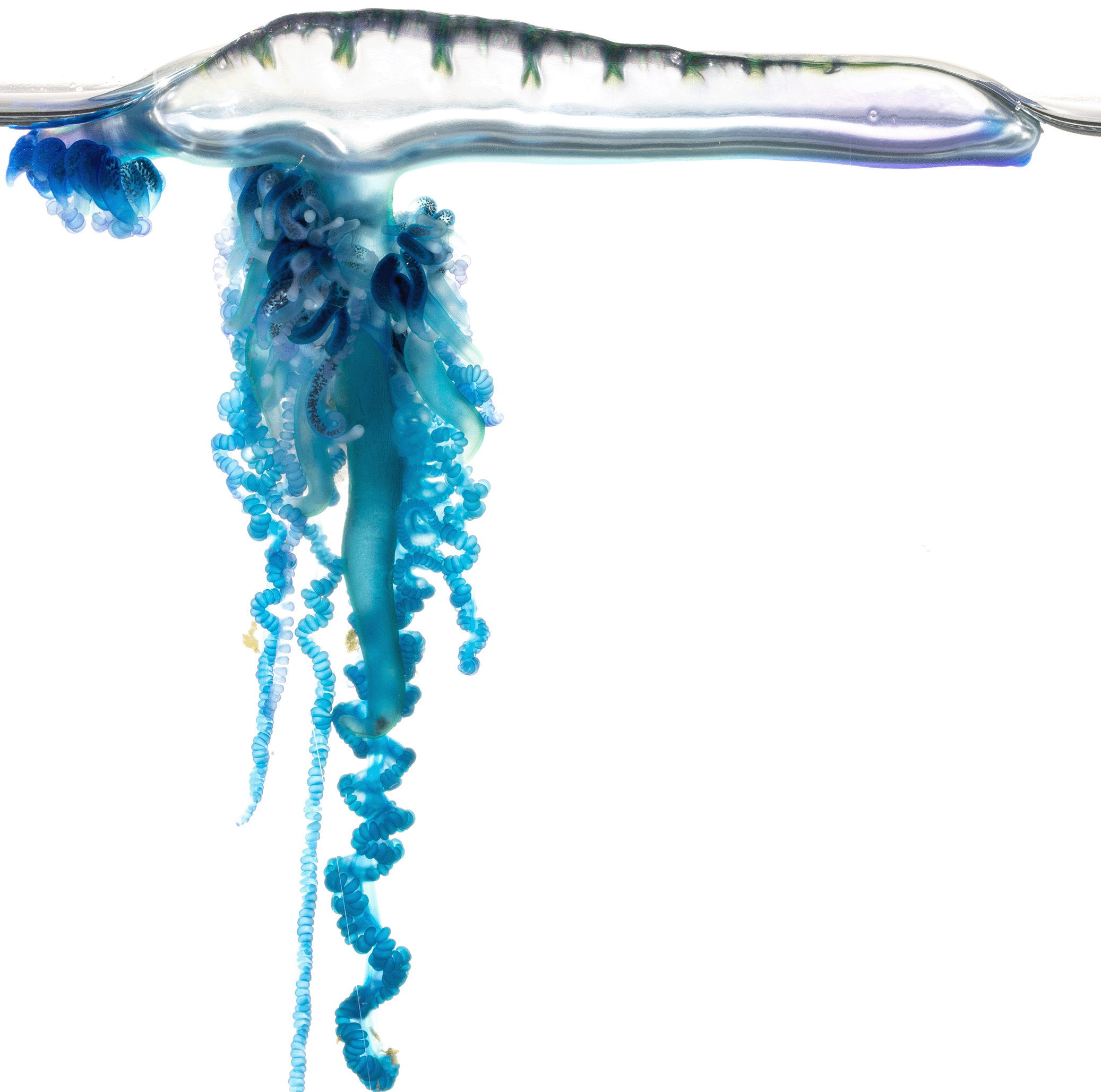
Sailing siphonophore jellyfish in the genus Physalia
Physalia jellyfish, known as Portuguese man o’ war or bluebottles, are beautiful, dangerous, and mysterious animals found at the ocean surface around the globe.
 Physalia megalista from New Zealand
Physalia megalista from New Zealand
Sailors, swimmers, and scientists have encountered these animals for centuries, and have learned a few important things: They are colonial, meaning they grow by adding more bodies via asexual budding. They have a muscular sail they use to travel. And they have a powerful, sometimes fatal, sting.
But there is much more we still don’t know, including when and where they reproduce, and how far they actually migrate. We led a global initiative to collect Physalia from the world’s oceans and found evidence for cryptic diversity, as well as regionally endemic subpopulations.
Life-history evolution and insect eggs
Life-history theory describes how traits like size, shape, and reproductive features are related across organisms, and over evolutionary time. We study how developmental biology can modulate these trait relationships.
In previous work, we curated a dataset of 10,000+ insect reproductive traits like egg size and shape, by digitizing records from the literature. Our team used this dataset to ask questions like:
- Which insects lay the biggest and smallest eggs?
- How is egg size related to the development of the embryo inside?
- What can the vast range of egg shapes and sizes tell us about the evolutionary process?
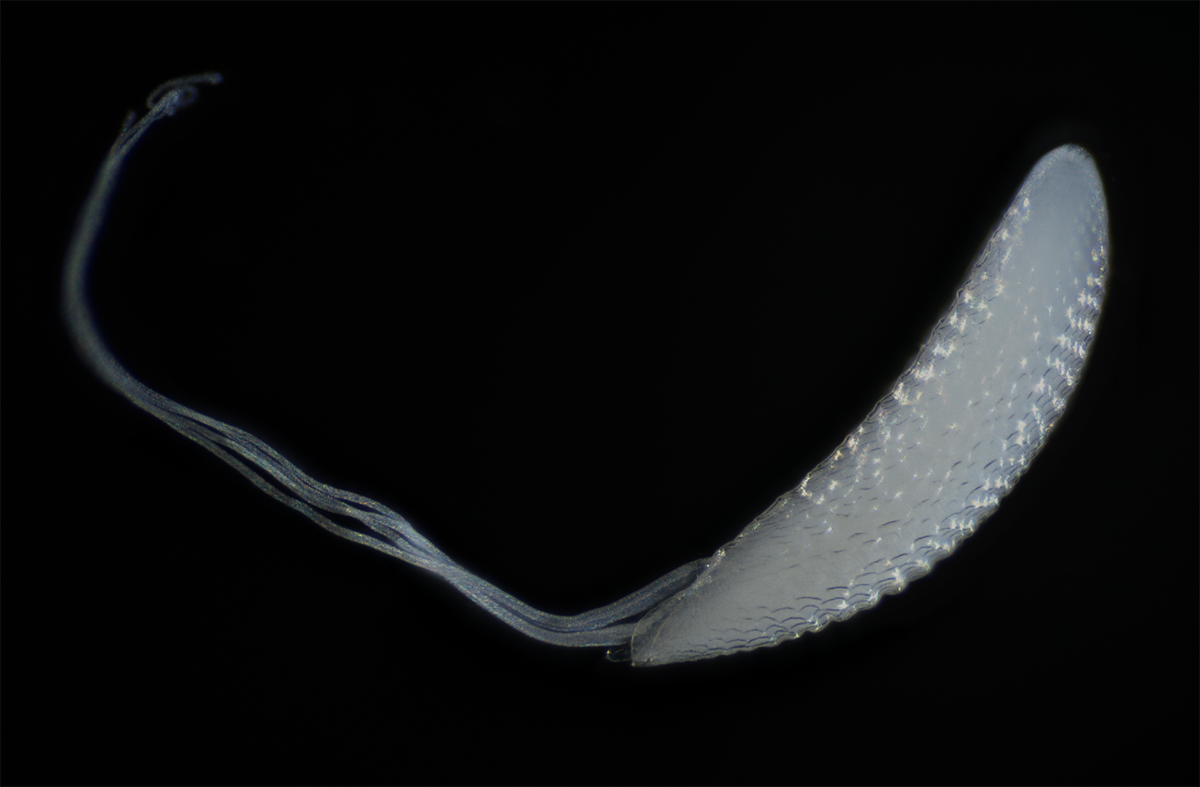 Egg of the Hawaiian fly Drosophila picticornis
Egg of the Hawaiian fly Drosophila picticornis
This work was published in the journal Nature. For a deep dive on insect egg shape and size, check out our custom data visualization
We also study the organ that makes insect eggs: the insect ovary. Our publication in Proceedings of the Royal Society B found that ovary development has a big impact on patterns in ovary structure variation across species.
Methods for evolutionary analysis of sequencing data
We develop open-source software and computational methods for the evolutionary analysis of sequencing datasets, spanning the full workflow from raw data processing through comparative phylogenetic inference.
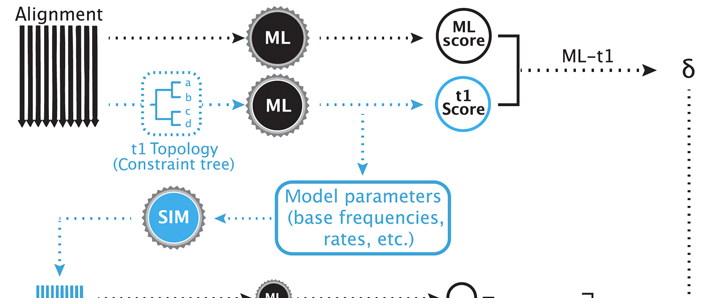
sowhat (GitHub) statistically tests competing phylogenetic hypotheses and has been applied to our own work on the evolutionary history of invertebrate lineages (e.g. Cnidaria, Siphonophora).
countland (GitHub) provides tools for evolutionary analysis of single-cell RNA-seq data using count-based approaches that avoid normalization assumptions.
sharkmer (GitHub) enables rapid, reference-free assembly of gene products directly from raw sequencing reads.
Our latest work uses AI to generate structured, queryable datasets from the published taxonomic literature, enabling richer biodiversity datasets for comparative evolutionary analysis.
Outreach
We are committed to a research approach that is open, inclusive, and accessible. If you are interested in collaborating in this endeavor, please reach out.
Samuel Church has a background in adult education, directing programs in New Haven, CT and Cambridge, MA, primarily serving Portuguese and Spanish-speaking communities.
People



GitHub





Contact
email: samuel.church@nyu.edu
office: Brown Building 24 Waverly Place, 9th Floor New York, NY 10003, USA